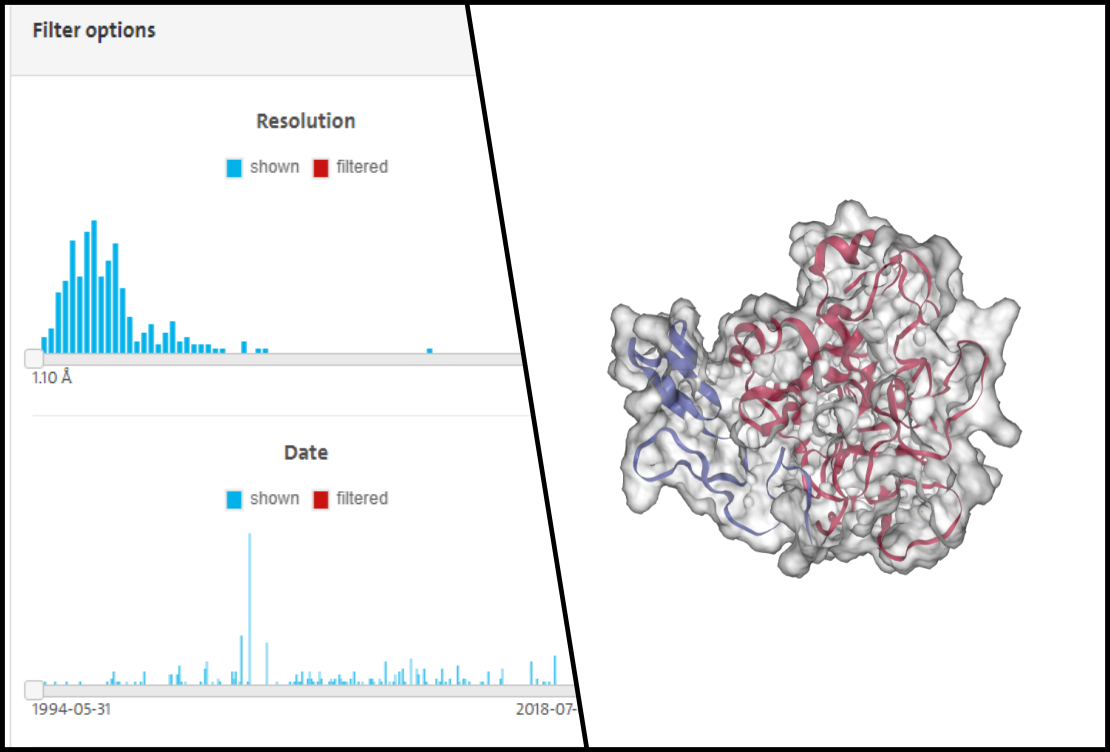
Tutorial 1: Getting started with Proteins Plus
This video will teach you some basics skills for working with Proteins Plus. You will learn how find to a crystal structure for your desired protein and then visualize it in different ways. 6 mins
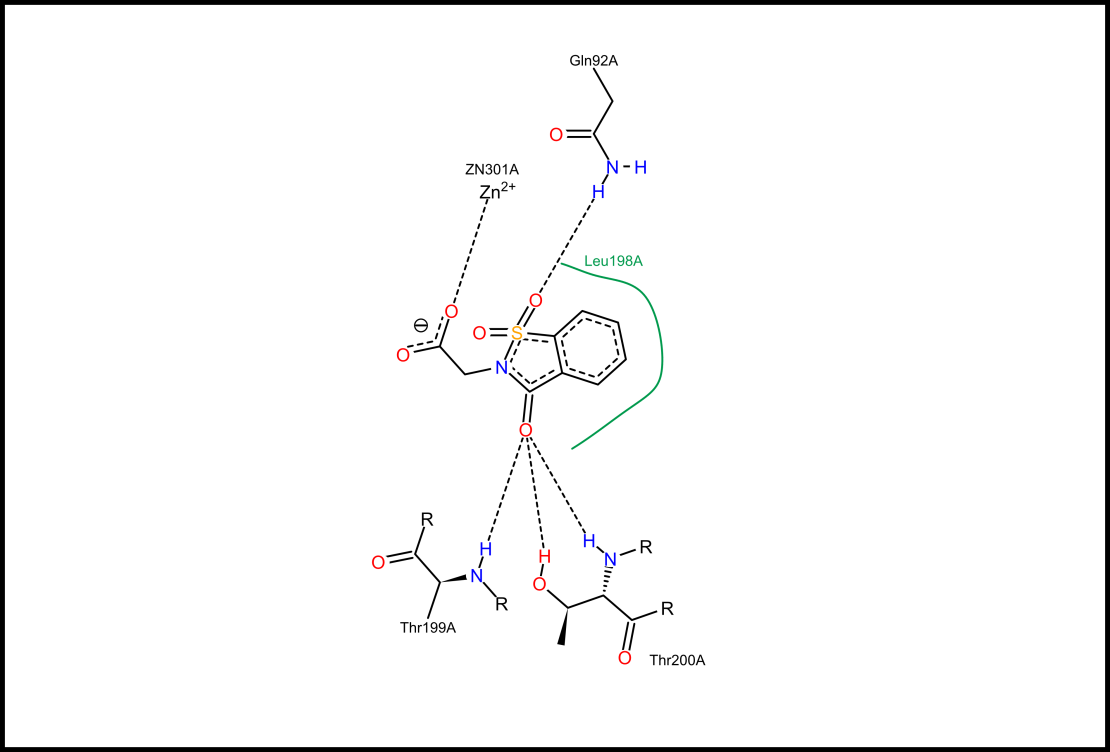
Tutorial 2: How does my protein interact with its ligands?
Understanding the interactions between proteins and their ligands is crucial for understanding the protein functionality and a first step in computational drug design. In this video you will learn how to identify and analyze these interactions with the tools supplied by Proteins Plus. 7 mins
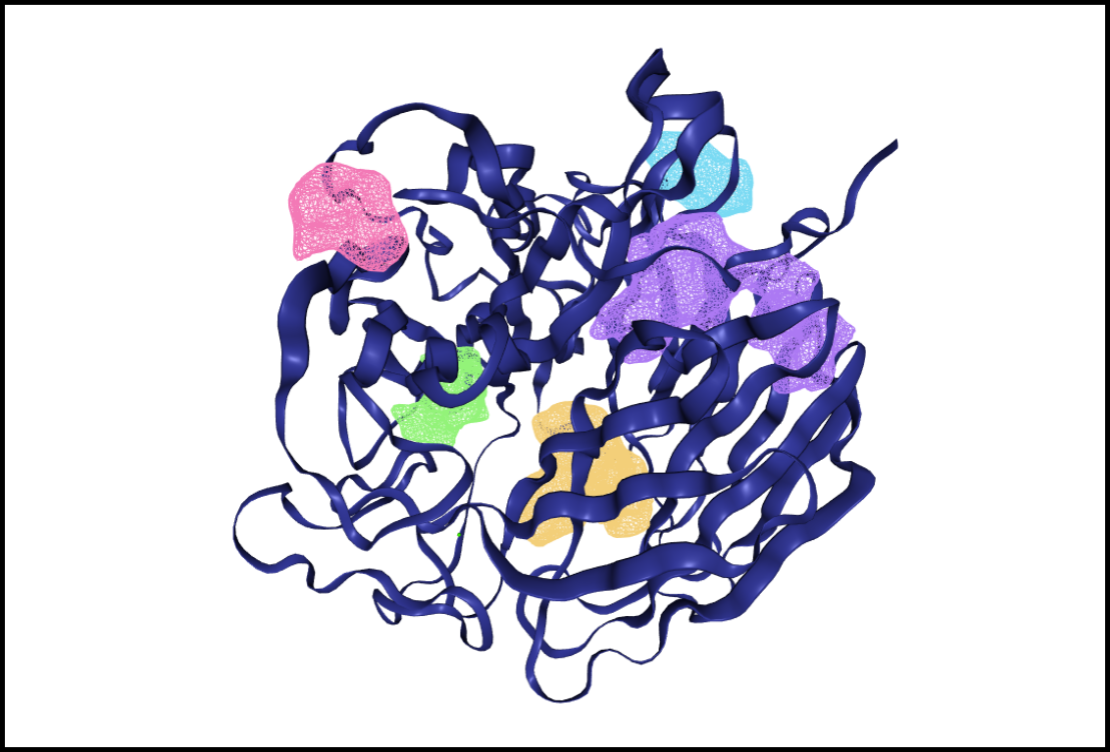
Tutorial 3: Where do I find druggable binding sites in my protein?
In this video you will learn how to identify binding sites in a crystal structure and estimate their druggability. Furthermore, you will compare the binding sites to similar pockets in the PDB to find potential ligands. 7 mins
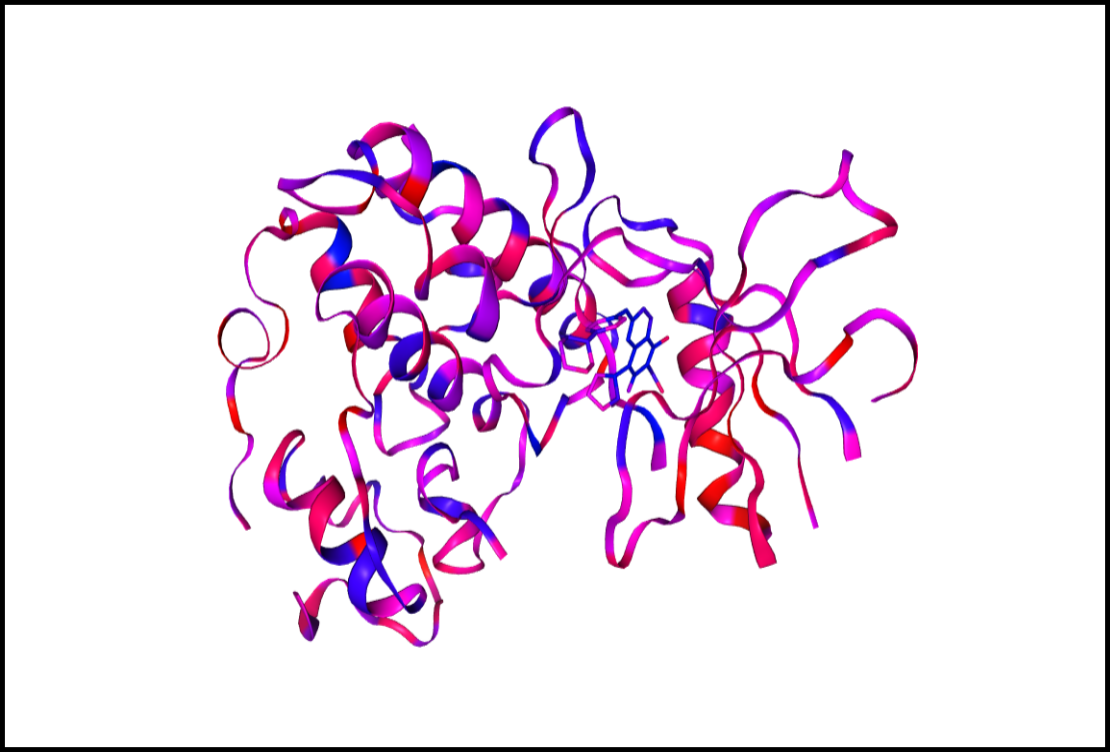
Tutorial 4: Is my crystal structure suitable for docking?
The success of virtual screening methods critically relies on the crystal structure used. In this video you will learn how to assess whether the quality of your crystal structure is sufficient for virtual screening using different Proteins Plus tools. 7 mins
Tutorial 5: Am I looking at a protein-protein interaction or just at a crystal artifact?
Protein-protein interactions (PPIs) form the basis for regulatory pathways in a cell and are thus involved in all major biological processes like metabolism, cell division or apoptosis. In this video you will learn how to discriminate PPIs from crystal artifacts. 4 mins


